This workflow uses Braker3 to annotate a genome.
Associated Tutorial
This workflows is part of the tutorial Genome annotation with Braker3, available in the GTN
Features
- Includes Galaxy Workflow Tests
- Includes a [Galaxy Workflow ...
Type: Nextflow
Creators: Peter J Bailey, Bailey PJ, Alexander Peltzer, Botvinnik O, Olga Botvinnik, Marques de Almeida F, Peltzer A, Sturm G
Submitter: WorkflowHub Bot
Alternative splicing analysis using RNA-seq.
Nextflow rnafusion analysis pipeline, part of the nf-core community.
Dual RNA-seq pipeline
Differential abundance analysis

MultiAffinity enables the study of how gene dysregulation propagates on a multilayer network on a disease of interest, uncovering key genes. Find the detailed documentation for the tool here.

Type: Common Workflow Language
Creators: Laura Rodriguez-Navas, Mar Batlle
Submitter: Laura Rodriguez-Navas
Flashlite-Trinity contains two workflows that run Trinity on the University of Queensland's HPC, Flashlite. Trinity performs de novo transcriptome assembly of RNA-seq data by combining three independent software modules Inchworm, Chrysalis and Butterfly to process RNA-seq reads. The algorithm can detect isoforms, handle paired-end reads, multiple insert sizes and strandedness. Users can run Flashlite-Trinity on single samples, or smaller samples requiring <500Gb ...
Type: Shell Script
Creators: Tracy Chew, Rosemarie Sadsad, Georgina Samaha, Cali Willet
Submitter: Tracy Chew
RNASeq-DE @ NCI-Gadi processes RNA sequencing data (single, paired and/or multiplexed) for differential expression (raw FASTQ to counts). This pipeline consists of multiple stages and is designed for the National Computational Infrastructure's (NCI) Gadi supercompter, leveraging multiple nodes to run each stage in parallel.
Infrastructure_deployment_metadata: Gadi (NCI)
Description: Trinity @ NCI-Gadi contains a staged Trinity workflow that can be run on the National Computational Infrastructure’s (NCI) Gadi supercomputer. Trinity performs de novo transcriptome assembly of RNA-seq data by combining three independent software modules Inchworm, Chrysalis and Butterfly to process RNA-seq reads. The algorithm can detect isoforms, handle paired-end reads, multiple insert sizes and strandedness. ...
Type: Shell Script
Creators: Georgina Samaha, Rosemarie Sadsad, Tracy Chew, Matthew Downton, Andrey Bliznyuk, Rika Kobayashi, Ben Menadue, Ben Evans
Submitter: Tracy Chew
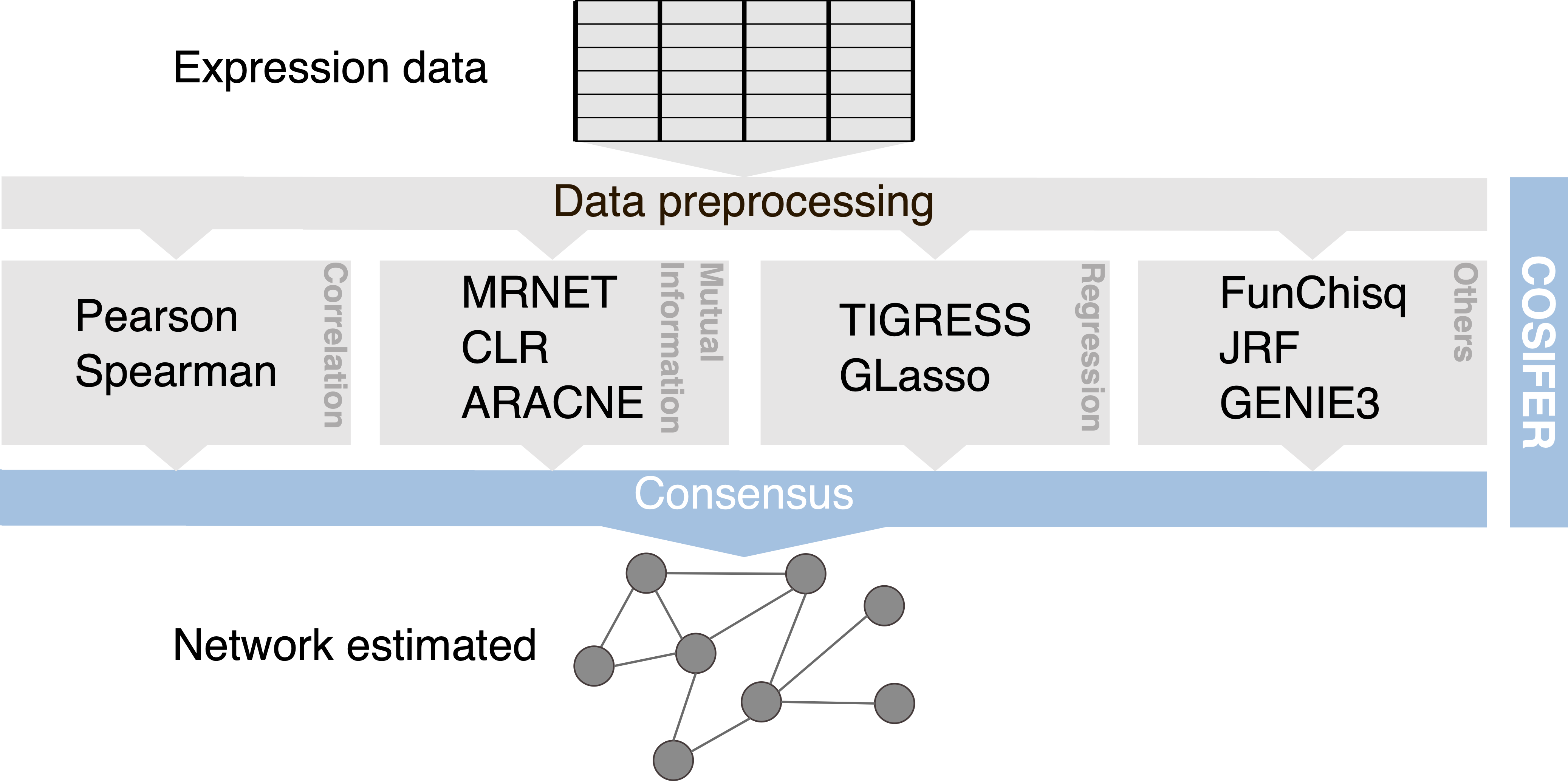
COnSensus Interaction Network InFErence Service
Inference framework for reconstructing networks using a consensus approach between multiple methods and data sources.

Reference
[Manica, Matteo, Charlotte, Bunne, Roland, Mathis, Joris, Cadow, Mehmet Eren, Ahsen, Gustavo A, Stolovitzky, and María Rodríguez, Martínez. "COSIFER: a python package for the consensus inference of molecular interaction ...
Type: Common Workflow Language
Creators: Laura Rodriguez-Navas, José Mª Fernández
Submitter: Laura Rodriguez-Navas
COnSensus Interaction Network InFErence Service
Inference framework for reconstructing networks using a consensus approach between multiple methods and data sources.
Reference
[Manica, Matteo, Charlotte, Bunne, Roland, Mathis, Joris, Cadow, Mehmet Eren, Ahsen, Gustavo A, Stolovitzky, and María Rodríguez, Martínez. "COSIFER: a python package for the consensus inference of molecular ...
Workflow for Spliced RNAseq data Steps:
- workflow_quality.cwl:
- FastQC (Read Quality Control)
- fastp (Read Trimming)
- STAR (Read mapping)
- featurecounts (transcript read counts)
- kallisto (transcript [pseudo]counts)
RNA-Seq pipeline
Here we provide the tools to perform paired end or single read RNA-Seq analysis including raw data quality control, differential expression (DE) analysis and functional annotation. As input files you may use either zipped fastq-files (.fastq.gz) or mapped read data (.bam files). In case of paired end reads, corresponding fastq files should be named using .R1.fastq.gz and .R2.fastq.gz suffixes.
Pipeline Workflow
All analysis steps are illustrated in the pipeline ...
 Tests All failing
Tests All failing




 Download
Download